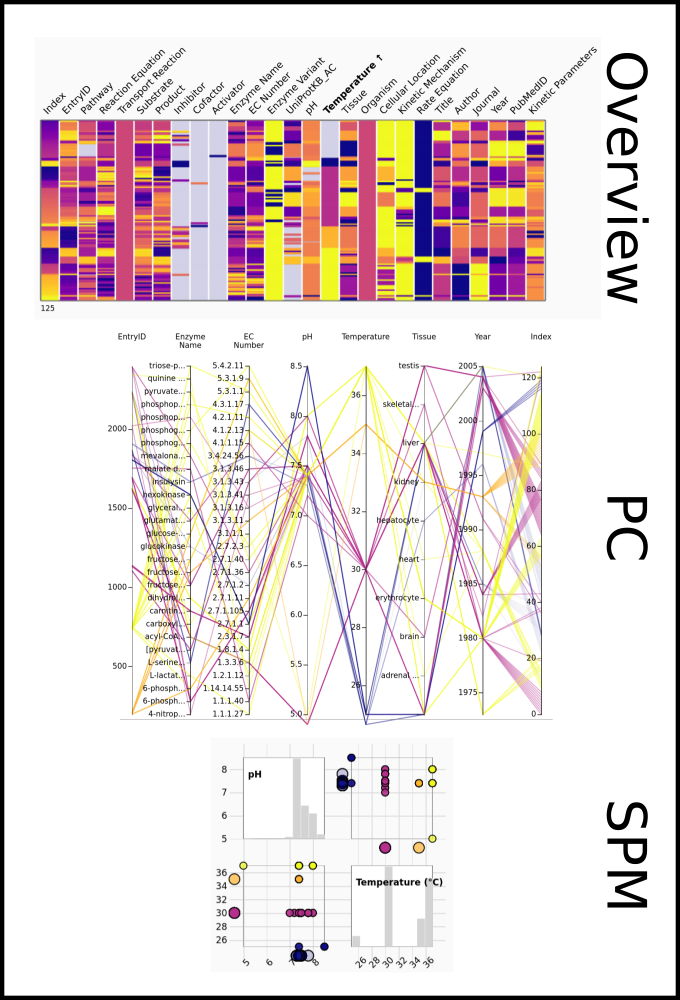
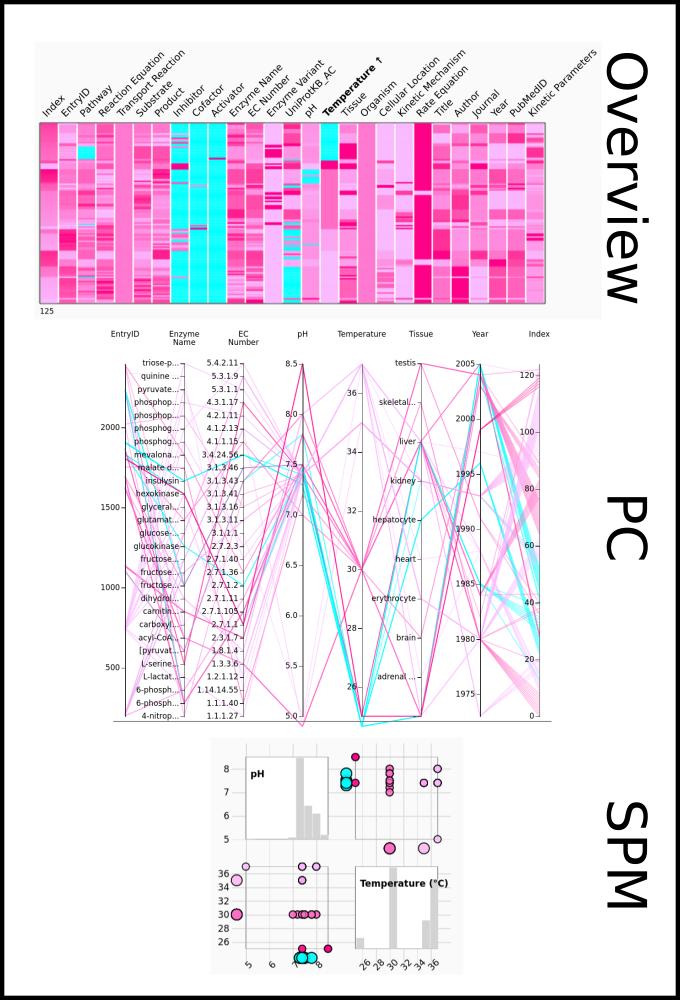
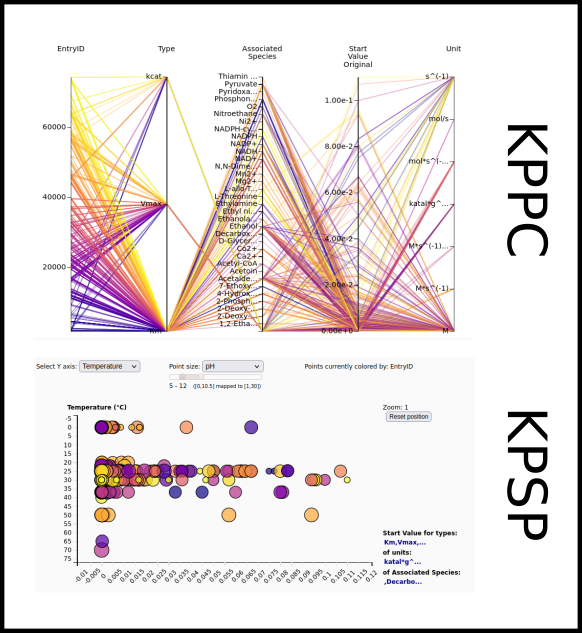
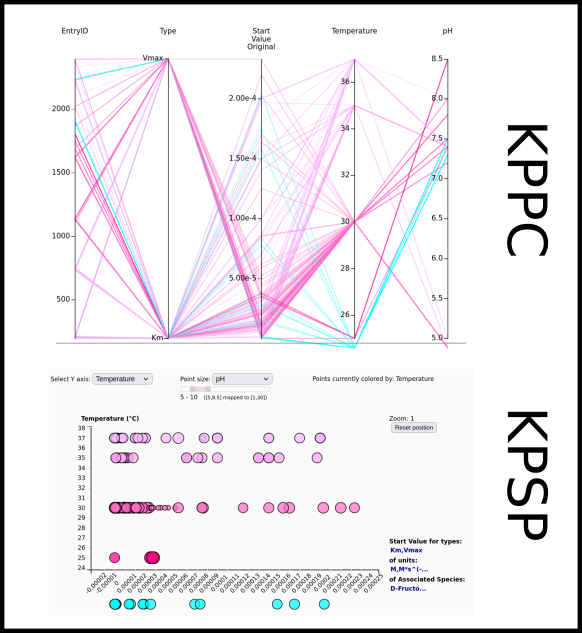
To improve SABIO-RK data interoperability, semantic markup was added to web pages as described and defined by
This structured information makes it easier to discover, collate and analyse our data.
|
74001
|
Purification and characterization of crotonase from Clostridium acetobutylicum.
|
|
74002
|
Purification and characterization of crotonase from Clostridium acetobutylicum.
|
|
74003
|
Catabolic alanine racemase from Salmonella typhimurium: DNA sequence, enzyme purification, and ...
|
|
74004
|
Catabolic alanine racemase from Salmonella typhimurium: DNA sequence, enzyme purification, and ...
|
|
74005
|
Purification and properties of diaminopimelic acid epimerase from Escherichia coli.
|
|
74006
|
Purification and properties of diaminopimelic acid epimerase from Escherichia coli.
|
|
74007
|
Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy
|
|
74008
|
Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy
|
|
74009
|
Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy
|
|
74010
|
Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy
|
|
74011
|
Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy
|
|
74012
|
Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy
|
|
74013
|
Vibrio cholerae FabV defines a new class of enoyl-acyl carrier protein reductase
|
|
74014
|
Vibrio cholerae FabV defines a new class of enoyl-acyl carrier protein reductase
|
|
74015
|
Vibrio cholerae FabV defines a new class of enoyl-acyl carrier protein reductase
|
|
74016
|
Vibrio cholerae FabV defines a new class of enoyl-acyl carrier protein reductase
|
|
74017
|
Vibrio cholerae FabV defines a new class of enoyl-acyl carrier protein reductase
|
|
74018
|
Vibrio cholerae FabV defines a new class of enoyl-acyl carrier protein reductase
|
|
74019
|
A class I fructose-1,6-bisphosphate aldolase is associated with salt stress tolerance in a halotolerant ...
|
|
74020
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74021
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74022
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74023
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74024
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74025
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74026
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74027
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74028
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74029
|
Structure determination of T cell protein-tyrosine phosphatase
|
|
74030
|
Diacylglycerol kinase from suspension cultured plant cells : purification and properties.
|
|
74031
|
Diacylglycerol kinase from suspension cultured plant cells : purification and properties.
|
|
74032
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74033
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74034
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74035
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74036
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74037
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74038
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74039
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74040
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74041
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74042
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74043
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74044
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74045
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74046
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74047
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74048
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74049
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74050
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74051
|
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase
|
|
74052
|
Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate ...
|
|
74053
|
Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate ...
|
|
74054
|
Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate ...
|
|
74055
|
Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate ...
|
|
74056
|
Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate ...
|
|
74057
|
Osmotic and chill activation of glycine betaine porter II in Listeria monocytogenes membrane vesicles
|
|
74058
|
Osmotic and chill activation of glycine betaine porter II in Listeria monocytogenes membrane vesicles
|
|
74059
|
Osmotic and chill activation of glycine betaine porter II in Listeria monocytogenes membrane vesicles
|
|
74060
|
Protein tyrosine phosphatase RQ is a phosphatidylinositol phosphatase that can regulate cell survival and ...
|
|
74061
|
Immobilization of thermostable exo-inulinase from mutant thermophilic Aspergillus tamarii-U4 using ...
|
|
74062
|
Immobilization of thermostable exo-inulinase from mutant thermophilic Aspergillus tamarii-U4 using ...
|
|
74063
|
Improvement of enzymological properties of pepsin by chemical modification with chitooligosaccharides
|
|
74064
|
Improvement of enzymological properties of pepsin by chemical modification with chitooligosaccharides
|
|
74065
|
Characterization of Two Endo-β-1, 4-Xylanases from Myceliophthora thermophila and Their ...
|
|
74066
|
Characterization of Two Endo-β-1, 4-Xylanases from Myceliophthora thermophila and Their ...
|
|
74067
|
Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway
|
|
74068
|
Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway
|
|
74069
|
Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway
|
|
74070
|
Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway
|
|
74071
|
Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway
|
|
74072
|
Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway
|
|
74073
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74074
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74075
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74076
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74077
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74078
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74079
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74080
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74081
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74082
|
Correlated Mutation in the Evolution of Catalysis in Uracil DNA Glycosylase Superfamily
|
|
74083
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74084
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74085
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74086
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74087
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74088
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74089
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74090
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74091
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74092
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74093
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74094
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74095
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74096
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74097
|
Aldose Reductase Differential Inhibitors in Green Tea
|
|
74098
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74099
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74100
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74101
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74102
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74103
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74104
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74105
|
Tyrosine aminotransferase catalyzes the final step of methionine recycling in Klebsiella pneumoniae
|
|
74106
|
Functional expression in Escherichia coli of low-affinity and high-affinity Na(+)(Li(+))/H(+) antiporters of ...
|
|
74107
|
Functional expression in Escherichia coli of low-affinity and high-affinity Na(+)(Li(+))/H(+) antiporters of ...
|
|
74108
|
Functional expression in Escherichia coli of low-affinity and high-affinity Na(+)(Li(+))/H(+) antiporters of ...
|
|
74109
|
Functional expression in Escherichia coli of low-affinity and high-affinity Na(+)(Li(+))/H(+) antiporters of ...
|
|
74110
|
Functional expression in Escherichia coli of low-affinity and high-affinity Na(+)(Li(+))/H(+) antiporters of ...
|
|
74111
|
Functional expression in Escherichia coli of low-affinity and high-affinity Na(+)(Li(+))/H(+) antiporters of ...
|
|
74112
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74113
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74114
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74115
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74116
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74117
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74118
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74119
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74120
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74121
|
Binding interaction of quercetin-3-beta-galactoside and its synthetic derivatives with SARS-CoV 3CL(pro): ...
|
|
74122
|
Recent advances in research on the most novel carbonic anhydrases, CA XIII and XV
|
|
74123
|
Recent advances in research on the most novel carbonic anhydrases, CA XIII and XV
|
|
74124
|
Recent advances in research on the most novel carbonic anhydrases, CA XIII and XV
|
|
74125
|
Recent advances in research on the most novel carbonic anhydrases, CA XIII and XV
|
|
74126
|
Yct1p, a novel, high-affinity, cysteine-specific transporter from the yeast Saccharomyces cerevisiae
|
|
74127
|
Purification and characterization of recombinant human cyclooxygenase-2.
|
|
74128
|
Purification and characterization of recombinant human cyclooxygenase-2.
|
|
74129
|
Structure- and function-based characterization of a new phosphoglycolate phosphatase from Thermoplasma ...
|
|
74130
|
Structure- and function-based characterization of a new phosphoglycolate phosphatase from Thermoplasma ...
|
|
74131
|
Structure- and function-based characterization of a new phosphoglycolate phosphatase from Thermoplasma ...
|
|
74132
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74133
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74134
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74135
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74136
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74137
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74138
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74139
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74140
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74141
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74142
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74143
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74144
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74145
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74146
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74147
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74148
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74149
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74150
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74151
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74152
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74153
|
Binuclear metal centers in plant purple acid phosphatases: Fe-Mn in sweet potato and Fe-Zn in soybean
|
|
74154
|
Identification and Biochemical Characterization of a Novel Protein Phosphatase 2C-Like Ser/Thr Phosphatase in ...
|
|
74155
|
Identification and Biochemical Characterization of a Novel Protein Phosphatase 2C-Like Ser/Thr Phosphatase in ...
|
|
74156
|
The crystal structure of three site-directed mutants of Escherichia coli dihydrodipicolinate synthase: further ...
|
|
74157
|
The crystal structure of three site-directed mutants of Escherichia coli dihydrodipicolinate synthase: further ...
|
|
74158
|
The crystal structure of three site-directed mutants of Escherichia coli dihydrodipicolinate synthase: further ...
|
|
74159
|
The crystal structure of three site-directed mutants of Escherichia coli dihydrodipicolinate synthase: further ...
|
|
74160
|
Peptidyl-alpha-hydroxyglycine alpha-amidating lyase. Purification, characterization, and expression.
|
|
74161
|
Peptidyl-alpha-hydroxyglycine alpha-amidating lyase. Purification, characterization, and expression.
|
|
74162
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74163
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74164
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74165
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74166
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74167
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74168
|
Essential features of the catalytic core of peptidyl-alpha-hydroxyglycine alpha-amidating lyase
|
|
74169
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74170
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74171
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74172
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74173
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74174
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74175
|
Use of endoproteases to identify catalytic domains, linker regions, and functional interactions in soluble ...
|
|
74176
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74177
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74178
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74179
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74180
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74181
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74182
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74183
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74184
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74185
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74186
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74187
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74188
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74189
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74190
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74191
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74192
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74193
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74194
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74195
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74196
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74197
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74198
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74199
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74200
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74201
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74202
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74203
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74204
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74205
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74206
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74207
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74208
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74209
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74210
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74211
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74212
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74213
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74214
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74215
|
Carbonic anhydrase activators: activation of the beta-carbonic anhydrases from the pathogenic fungi Candida ...
|
|
74216
|
4-Hydroxybenzoate 3-geranyltransferase from Lithospermum erythrorhizon: purification of a plant membrane-bound ...
|
|
74217
|
4-Hydroxybenzoate 3-geranyltransferase from Lithospermum erythrorhizon: purification of a plant membrane-bound ...
|
|
74218
|
A fibrinogen-clotting serine proteinase from Cerastes cerastes (horned viper) venom with arginine-esterase and ...
|
|
74219
|
A fibrinogen-clotting serine proteinase from Cerastes cerastes (horned viper) venom with arginine-esterase and ...
|
|
74220
|
A fibrinogen-clotting serine proteinase from Cerastes cerastes (horned viper) venom with arginine-esterase and ...
|
|
74221
|
A fibrinogen-clotting serine proteinase from Cerastes cerastes (horned viper) venom with arginine-esterase and ...
|
|
74222
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74223
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74224
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74225
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74226
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74227
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74228
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74229
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74230
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74231
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74232
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74233
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74234
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74235
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74236
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74237
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74238
|
Differential redox and electron-transfer properties of purified yeast, plant and human NADPH-cytochrome P-450 ...
|
|
74239
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74240
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74241
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74242
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74243
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74244
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74245
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74246
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74247
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74248
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74249
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74250
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74251
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74252
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74253
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74254
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74255
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74256
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74257
|
Ring-substituted 8-hydroxyquinoline-2-carboxanilides as photosystem II inhibitors
|
|
74258
|
Three-dimensional structure of phosphoenolpyruvate carboxylase: a proposed mechanism for allosteric inhibition
|
|
74259
|
Three-dimensional structure of phosphoenolpyruvate carboxylase: a proposed mechanism for allosteric inhibition
|
|
74260
|
Three-dimensional structure of phosphoenolpyruvate carboxylase: a proposed mechanism for allosteric inhibition
|
|
74261
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74262
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74263
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74264
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74265
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74266
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74267
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74268
|
Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium ...
|
|
74269
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74270
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74271
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74272
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74273
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74274
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74275
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74276
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74277
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74278
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74279
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74280
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74281
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74282
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74283
|
Structure-based design and synthesis of highly potent SARS-CoV 3CL protease inhibitors.
|
|
74284
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74285
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74286
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74287
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74288
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74289
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74290
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74291
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74292
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74293
|
Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the ...
|
|
74294
|
Bromo-Cyclobutenaminones as New Covalent UDP-N-Acetylglucosamine Enolpyruvyl Transferase (MurA) ...
|
|
74295
|
Bromo-Cyclobutenaminones as New Covalent UDP-N-Acetylglucosamine Enolpyruvyl Transferase (MurA) ...
|
|
74296
|
Bromo-Cyclobutenaminones as New Covalent UDP-N-Acetylglucosamine Enolpyruvyl Transferase (MurA) ...
|
|
74297
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74298
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74299
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74300
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74301
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74302
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74303
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74304
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74305
|
Screening of compound library identifies novel inhibitors against the MurA enzyme of Escherichia coli
|
|
74306
|
Genetic resistance to purine nucleoside phosphorylase inhibition in Plasmodium falciparum
|
|
74307
|
Genetic resistance to purine nucleoside phosphorylase inhibition in Plasmodium falciparum
|
|
74308
|
Genetic resistance to purine nucleoside phosphorylase inhibition in Plasmodium falciparum
|
|
74309
|
Genetic resistance to purine nucleoside phosphorylase inhibition in Plasmodium falciparum
|
|
74310
|
Genetic resistance to purine nucleoside phosphorylase inhibition in Plasmodium falciparum
|
|
74311
|
Genetic resistance to purine nucleoside phosphorylase inhibition in Plasmodium falciparum
|
|
74312
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74313
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74314
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74315
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74316
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74317
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74318
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74319
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74320
|
Methylcobamide:coenzyme M methyltransferase isozymes from Methanosarcina barkeri. Physicochemical ...
|
|
74321
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74322
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74323
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74324
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74325
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74326
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74327
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74328
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74329
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74330
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74331
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74332
|
Functional analysis of Gln-237 mutants of HhaI methyltransferase.
|
|
74333
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74334
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74335
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74336
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74337
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74338
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74339
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74340
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74341
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74342
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74343
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74344
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74345
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74346
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74347
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74348
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74349
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74350
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74351
|
Abietane-Type Diterpenoids Inhibit Protein Tyrosine Phosphatases by Stabilizing an Inactive Enzyme ...
|
|
74352
|
Polysaccharide deacetylases serve as new targets for the design of inhibitors against Bacillus anthracis and ...
|
|
74353
|
Polysaccharide deacetylases serve as new targets for the design of inhibitors against Bacillus anthracis and ...
|
|
74354
|
Polysaccharide deacetylases serve as new targets for the design of inhibitors against Bacillus anthracis and ...
|
|
74355
|
Polysaccharide deacetylases serve as new targets for the design of inhibitors against Bacillus anthracis and ...
|
|
74356
|
Polysaccharide deacetylases serve as new targets for the design of inhibitors against Bacillus anthracis and ...
|
|
74357
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74358
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74359
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74360
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74361
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74362
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74363
|
Characterization of human nicotinate phosphoribosyltransferase: Kinetic studies, structure prediction and ...
|
|
74364
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74365
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74366
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74367
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74368
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74369
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74370
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74371
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74372
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74373
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74374
|
Simultaneous involvement of a tungsten-containing aldehyde:ferredoxin oxidoreductase and a phenylacetaldehyde ...
|
|
74375
|
On the two components of pyridoxal 5'-phosphate synthase from Bacillus subtilis.
|
|
74376
|
On the two components of pyridoxal 5'-phosphate synthase from Bacillus subtilis.
|
|
74377
|
On the two components of pyridoxal 5'-phosphate synthase from Bacillus subtilis.
|
|
74378
|
On the two components of pyridoxal 5'-phosphate synthase from Bacillus subtilis.
|
|
74379
|
Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate ...
|
|
74380
|
Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate ...
|
|
74381
|
Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate ...
|
|
74382
|
Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate ...
|
|
74383
|
Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate ...
|
|
74384
|
Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate ...
|
|
74385
|
Comparative analysis of ribosome-associated adenosinetriphosphatase (ATPase) from pig liver and the ATPase of ...
|
|
74386
|
Comparative analysis of ribosome-associated adenosinetriphosphatase (ATPase) from pig liver and the ATPase of ...
|
|
74387
|
Comparative analysis of ribosome-associated adenosinetriphosphatase (ATPase) from pig liver and the ATPase of ...
|
|
74388
|
Comparative analysis of ribosome-associated adenosinetriphosphatase (ATPase) from pig liver and the ATPase of ...
|
|
74389
|
Comparative analysis of ribosome-associated adenosinetriphosphatase (ATPase) from pig liver and the ATPase of ...
|
|
74390
|
Comparative analysis of ribosome-associated adenosinetriphosphatase (ATPase) from pig liver and the ATPase of ...
|
|
74391
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74392
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74393
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74394
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74395
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74396
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74397
|
Kinetic analysis of the 4-methylideneimidazole-5-one-containing tyrosine aminomutase in enediyne antitumor ...
|
|
74398
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74399
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74400
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74401
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74402
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74403
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74404
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74405
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74406
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74407
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74408
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74409
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74410
|
Characterization of theanine-forming enzyme from Methylovorus mays no. 9 in respect to utilization of theanine ...
|
|
74411
|
Discovery of a Fungal Multicopper Oxidase That Catalyzes the Regioselective Coupling of a Tricyclic ...
|
|
74412
|
Discovery of a Fungal Multicopper Oxidase That Catalyzes the Regioselective Coupling of a Tricyclic ...
|
|
74413
|
Mutations in SELENBP1, encoding a novel human methanethiol oxidase, cause extraoral halitosis
|
|
74414
|
Mutations in SELENBP1, encoding a novel human methanethiol oxidase, cause extraoral halitosis
|
|
74415
|
Mutations in SELENBP1, encoding a novel human methanethiol oxidase, cause extraoral halitosis
|
|
74416
|
Human erythrocyte histamine N-methyltransferase: radiochemical microassay and biochemical properties.
|
|
74417
|
Human erythrocyte histamine N-methyltransferase: radiochemical microassay and biochemical properties.
|
|
74418
|
Human erythrocyte histamine N-methyltransferase: radiochemical microassay and biochemical properties.
|
|
74419
|
Mouse histamine N-methyltransferase: cDNA cloning, expression, gene cloning and chromosomal localization
|
|
74420
|
Mouse histamine N-methyltransferase: cDNA cloning, expression, gene cloning and chromosomal localization
|





 If the current search query is a refinement of the previous query "Use Previous Query (Go Back)" button can be used to perform the previous query again. Only one automatic step back is possible, but the query can be manually manipulated if desired.
If the current search query is a refinement of the previous query "Use Previous Query (Go Back)" button can be used to perform the previous query again. Only one automatic step back is possible, but the query can be manually manipulated if desired.